
Cancer of Unknown Primary : Comprehensive Genomic Profiling
INTRODUCTION
-
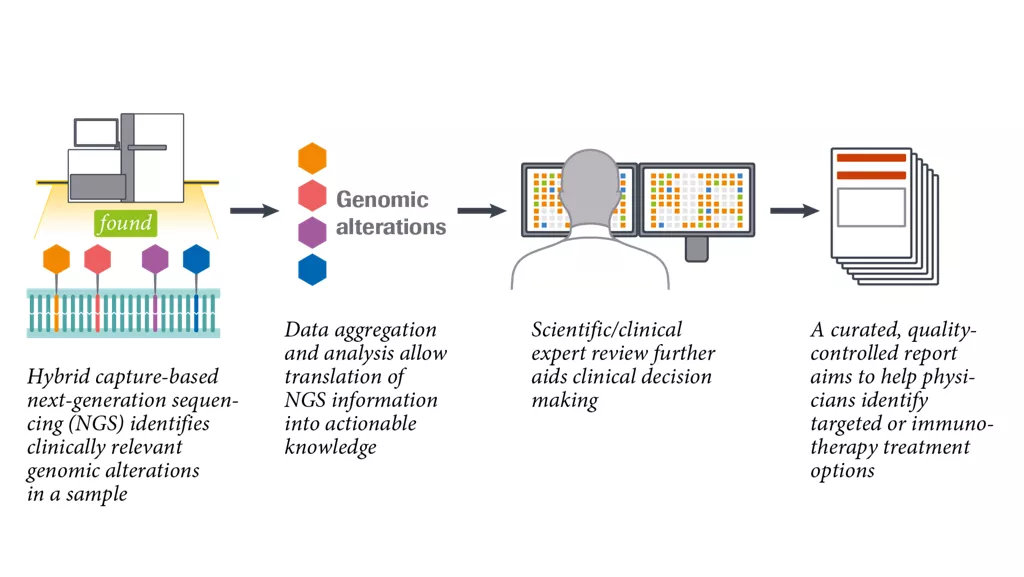
What is the comprehensive genomic profiling approach?
Comprehensive genomic profiling (CGP) can be conducted across cancer types to translate genomic data into clinically relevant information. It could help to inform molecularly-guided treatment options in CUP.1-4 -
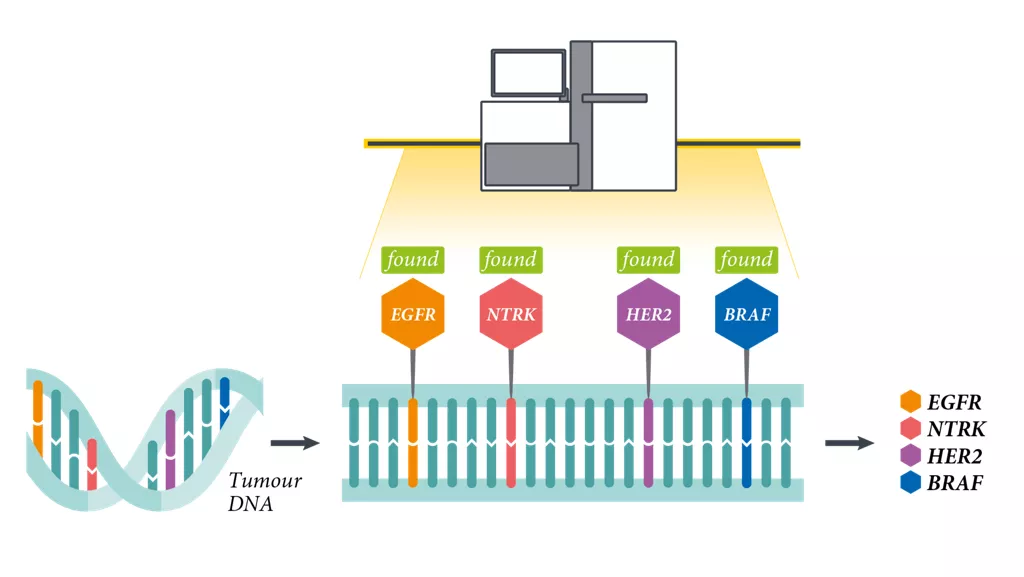
What does CGP detect?
CGP utilises next-generation sequencing (NGS) technology to detect both known and novel variants across the four main classes of genomic alterations in a large subset of cancer-related genes and identifies genomic signatures, i.e. tumour mutational burden (TMB) and microsatellite instability (MSI).4–9
-
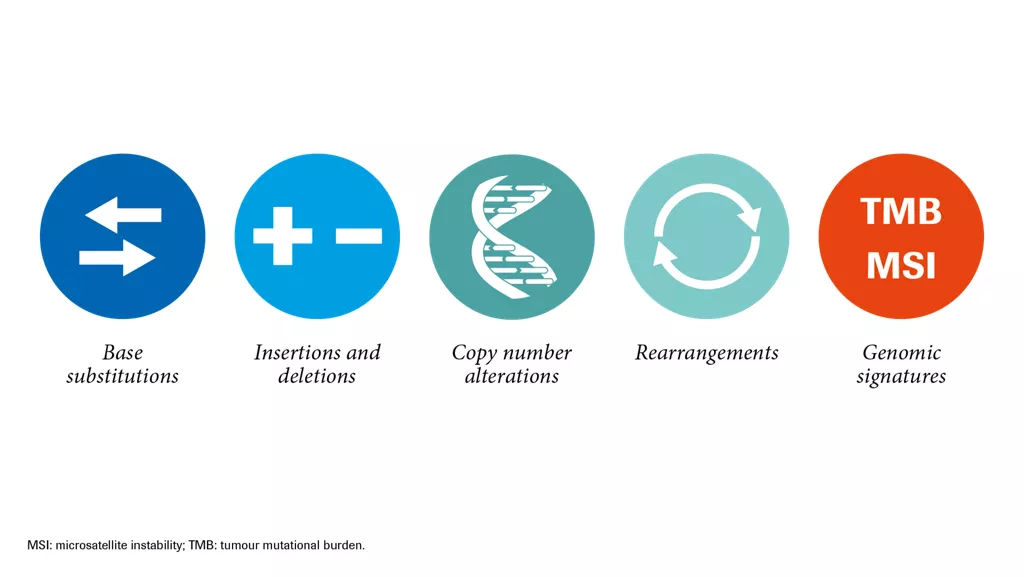
What types of alterations can CGP identify?
As an example, all Foundation Medicine services detect the four main classes of genomic alterations in a single test. FoundationOne®CDx additionally reports TMB and MSI, and FoundationOne®Liquid reports MSI.3-5,10
1. Ross JS et al. JAMA Oncol 2015; 1: 40–9.
2. Varghese AM et al. Ann Oncol 2017; 28: 3015–21.
3. Foundation Medicine, Inc.: Genomic Testing 2018.
Available at:
www.foundationmedicine.com/genomic-testing
(last accessed March 2019).
4. Frampton GM et al. Nat biotechnol 2013; 31: 1023–31.
5. He J et al. Blood 2016; 127: 3004–14.
6. Gagan J and van Allen EM. Genome Med 2015; 7: 80.
7. Rozenblum AB et al. J Thorac Oncol 2017; 12: 258–68.
8. Suh JH et al. Oncologist 2016; 21: 684–91.
9. NCCN Clinical Practice Guidelines in Oncology:
Non-Small Lung Cancer Version 6.2019. Available at: www.nccn.org/professionals/physician_gls/recently_updated.aspx
(last accessed January 2019).
10. Chalmers Z R et al. Genome Med 2017; 9: 34.
COMPREHENSIVE GENOMIC PROFILING VS GENE EXPRESSION PROFILING
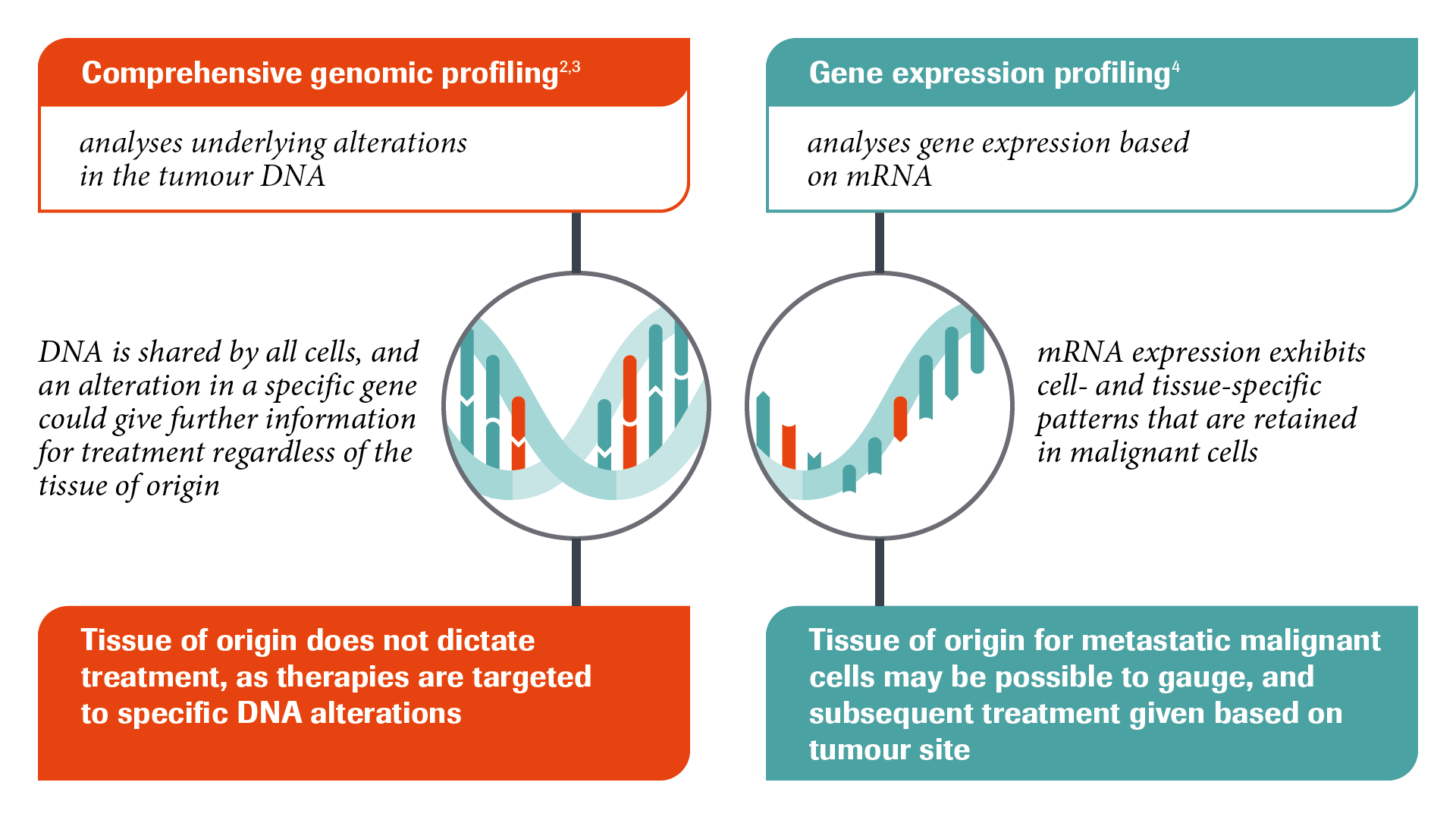
How do CGP and gene expression profiling (GEP) compare?
As mentioned, CGP identifies underlying alterations in the tumour DNA, whereas, GEP analyses gene expression patterns based on mRNA. Gene expression patterns can be recognised in a majority of metastases, where patterns may reflect their tissue of origin.1-4
1. Chalmers ZR et al. Genome Med 2017; 9: 34.
2. Frampton GM et al. Nat biotechnol 2013; 31: 1023–31.
3. He J et al. Blood 2016; 127: 3004–14.
4. Greco FA et al. Ann Oncol 2012; 23: 298–304.
AN EXAMPLE OF CGP'S UTILITY IN LUNG CANCER
CGP exemplified in non-small cell lung cancer (NSCLC)
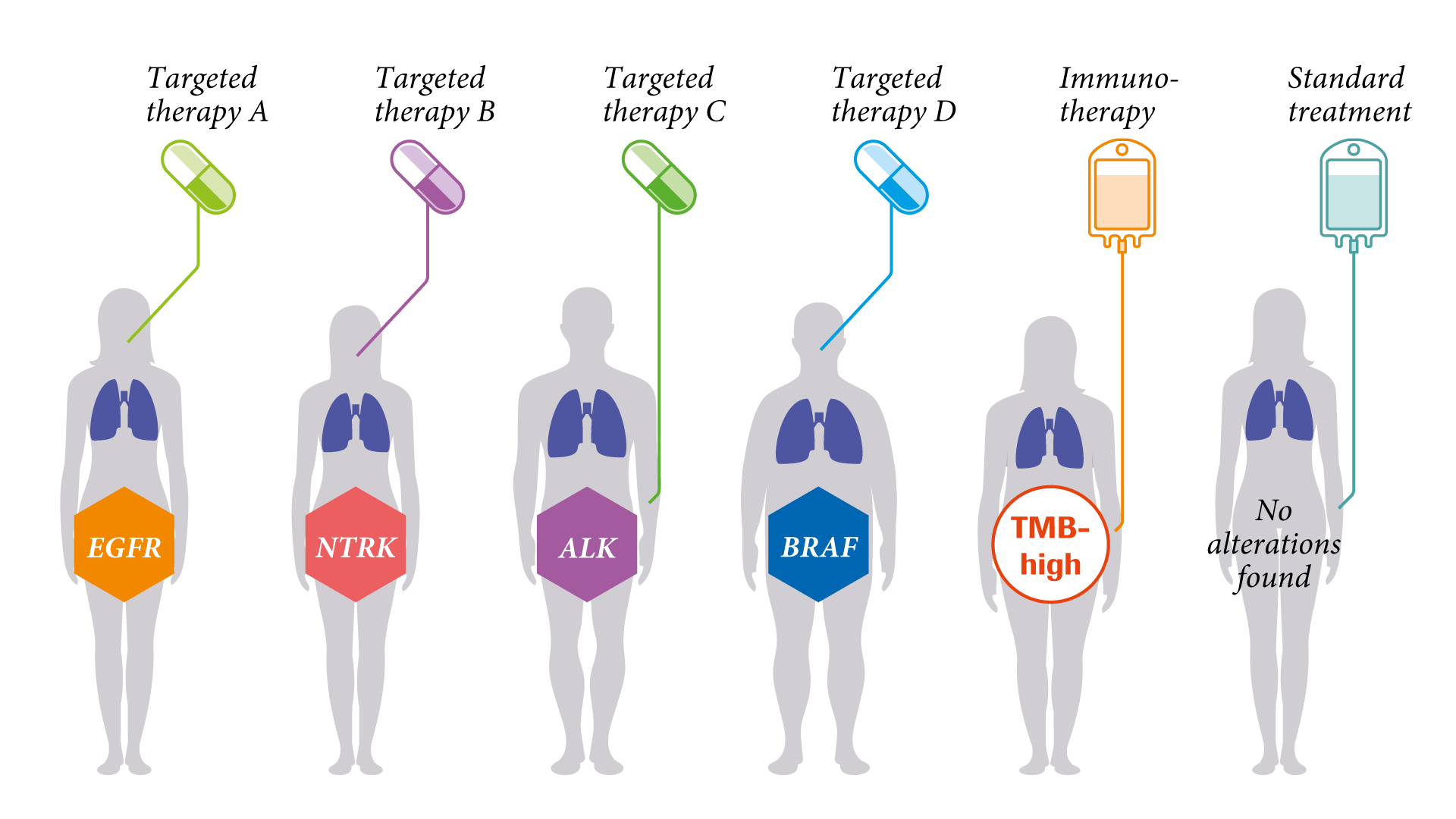
CGP, with validated multi-gene panel tests, identifies potentially clinically relevant genomic alterations in the tumour tissue. Treatment options can be tailored to the patient’s alteration pattern.1
At the forefront of this development is NSCLC, for which targeted therapies have been clinically validated and demonstrated to improve outcomes.2–8
NSCLC was thought to be one disease. We now know that many different alterations are driving lung cancer. Looking at the variety of NSCLC types, we realise that the variety of the genomic profiles reveals different entities. By utilising CGP, NSCLC discloses a variety of small molecular subtypes.
1. Foundation Medicine, Inc. 2018. Available at: www.foundationmedicine.com (last accessed March 2019).
2. Drilon A et al. N Engl J Med 2018; 378: 731–9.
3. Planchard D et al. Lancet Oncol 2016; 17: 984–93.
4. Hellmann MD et al. N Engl J Med 2018; 378: 2093–104.
5. Peters S et al. N Engl J Med 2017; 377: 829–38.
6. Soria JC et al. N Engl J Med 2018; 378: 113–25.
7. Rosell R et al. Lancet Oncol 2012; 13: 239–46.
8. Sequist LV et al. J Clin Oncol 2013; 31: 3327–34.
THE POTENTIAL OF CGP IN CUP SYNDROME
What is the potential of CGP in CUP Syndrome?
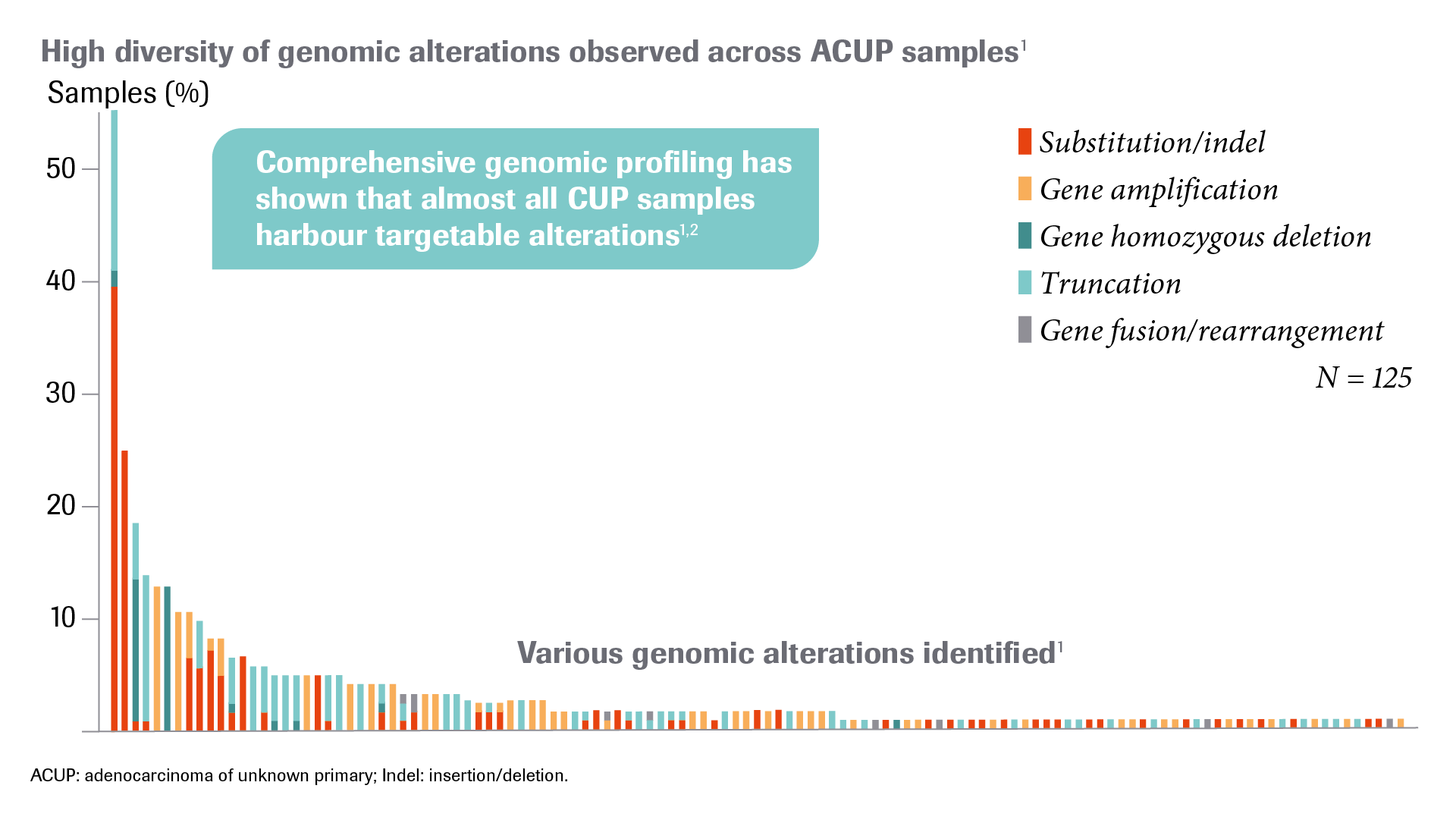
The majority of CUP patients (80–85%) have an unfavourable risk profile and currently have a poor prognosis despite treatment with different chemotherapy combinations in clinical trials. In this specific group, the treatment response rate is low and the median overall survival is generally less than one year. CGP has shown that almost all CUP samples harbour targetable alterations, potentially addressing the limited options and poor prognoses of patients with CUP.1,2
1. Ross JS et al. JAMA Oncol 2015; 1: 40–9.
2. Subbiah IM et al. Oncoscience 2017; 4: 47–56.
MOLECULARY-GUIDED THERAPY OPTIONS
CUP patients with poor prognosis receive standard therapy: platinum-based doublet chemotherapy or paclitaxel. In several phase II studies using platinum/taxane combinations or paclitaxel the overall response rates were approximately 30% and the median survival rates ranged from 7 to 11 months.1-3
Patients with CUP represent a heterogeneous population, harbouring a variety of potentially targetable alterations. One publication showed that 30% of patients had potentially targetable genomic alterations identified by molecular tumour profiling.4
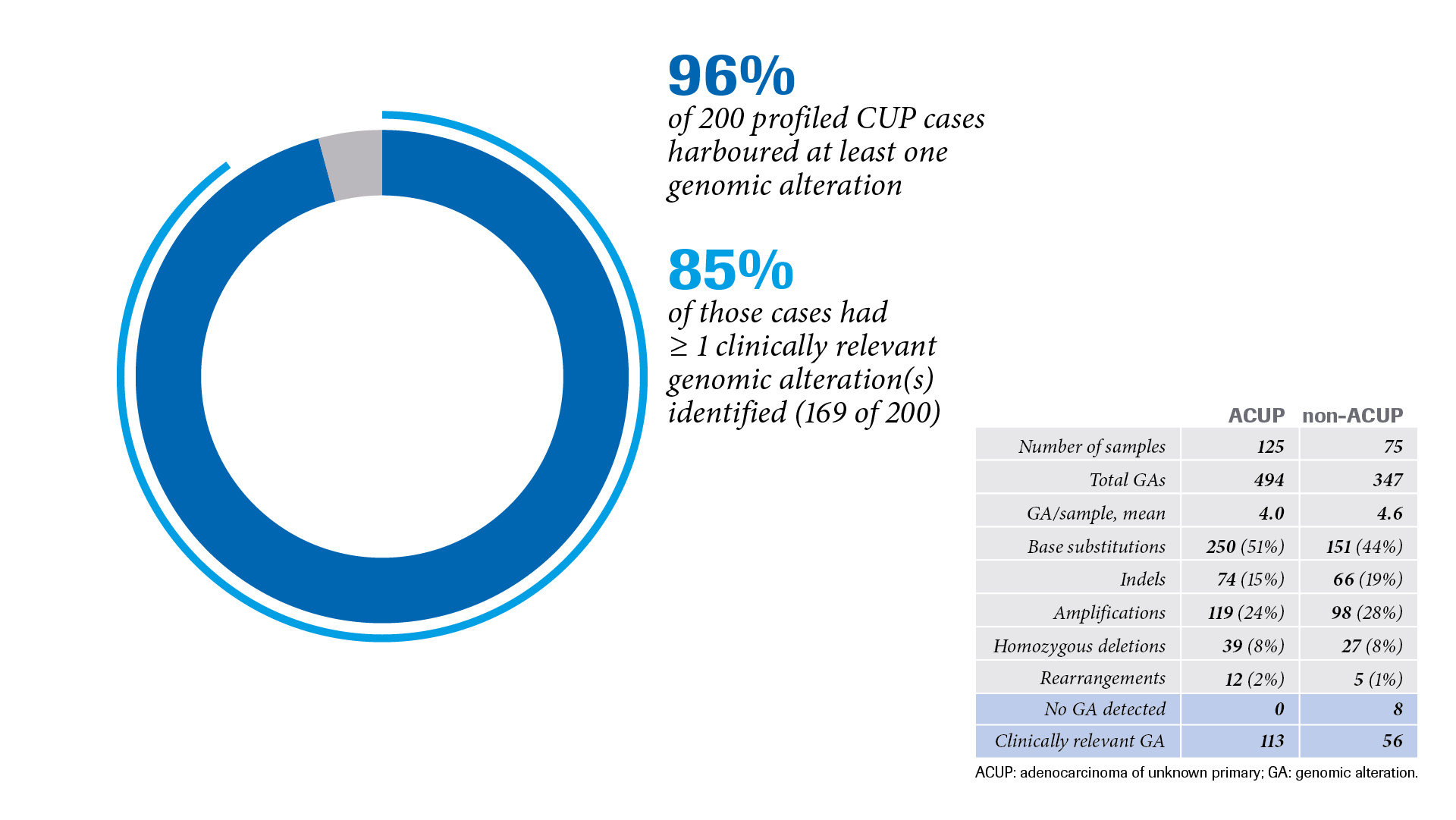
As described in another publication, samples from 200 CUP cases were analysed with CGP (Foundation Medicine® tissue biopsy assay) to determine whether it is possible to identify genomic alterations that can serve as a basis for targeted therapy in CUP.5 Of the 200 CUP cases, 96% exhibited at least one genomic alteration, with one or more clinically relevant genomic alterations identified in 85% of these cases (169 of 200).5
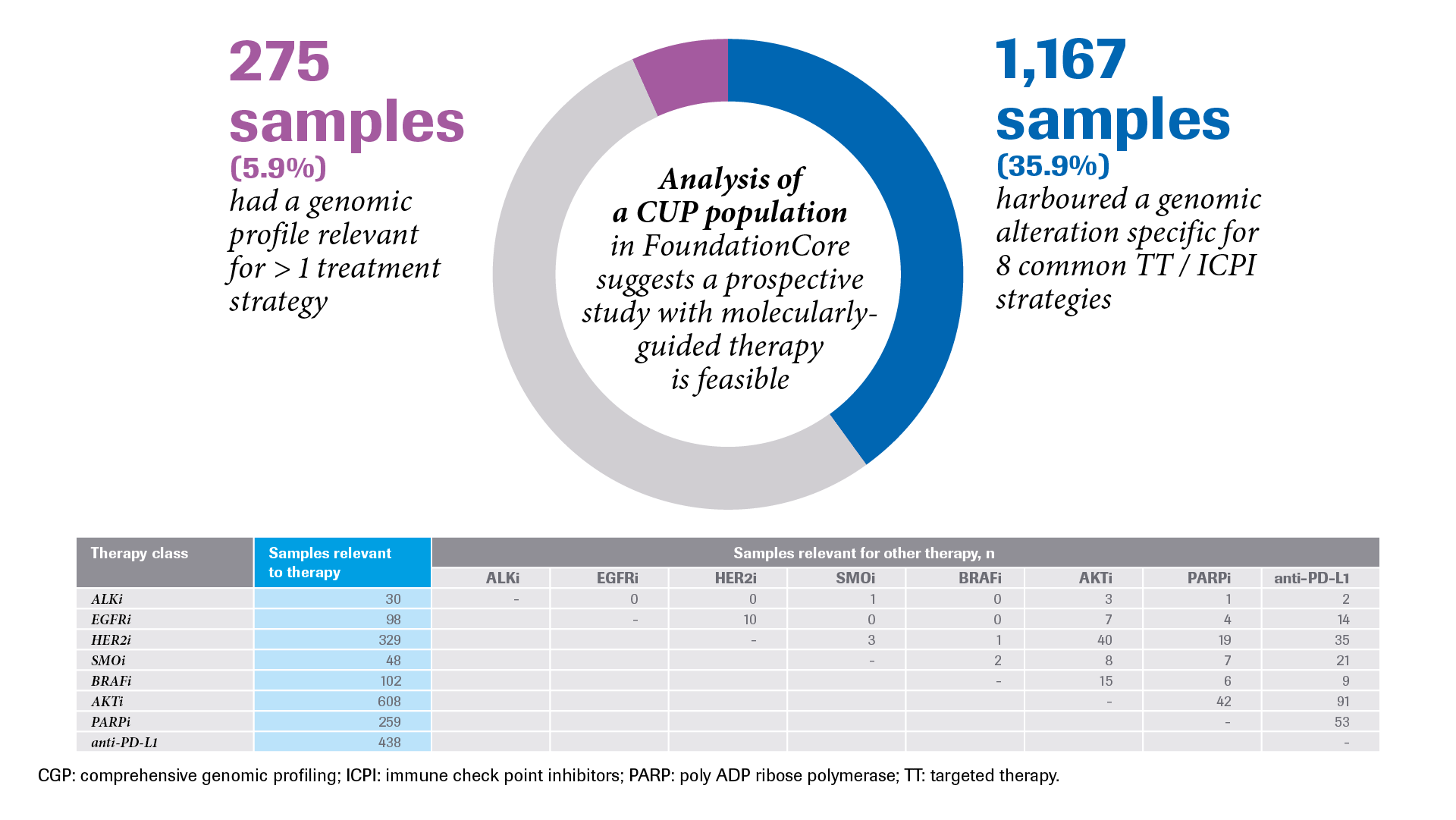
Analysis of CUP patient data from the FoundationCoreTM database (consisting of Foundation Medicine data) also indicated that a prospective clinical trial with molecularly-guided therapies is clinically feasible.6
1. Huebner G et al. British Journal of Cancer 2009; 100: 44–9.
2. Hainsworth JD et al. J Clin Oncol 1997; 15: 2385–93.
3. Greco FA et al. J Clin Oncol 2002; 15 (20): 1651–6.
4. Varghese AM et al. Ann Oncol 2017; 1 (28): 3015–21.
5. Ross JS et al. JAMA Oncol 2015; 1: 40–9.
6. Krämer A et al. J Clin Oncol 2018; 36 (15): (suppl e24162).
